Complex microbial communities within the human body, constituting the microbiome, have a broad impact on human health and disease. A growing body of research now examines the role of the microbiome in patients with critical illness, such as sepsis and acute respiratory failure. In this article, we provide an introduction to microbiome concepts and terminology and we systematically review the current evidence base of the critical-illness microbiome, including 51 studies in animal models and pediatric and adult critically-ill patients. We further examine how this emerging scientific discipline may transform the way we manage infectious and inflammatory diseases in intensive care units. The evolving molecular, culture-independent techniques offer the ability to study microbial communities in unprecedented depth and detail, and in the short-term, may enable us to diagnose and treat infections in critical care more precisely and effectively. Longer-term, these tools may also give us insights in the underlying pathophysiology of critical illness and reveal previously unsuspected targets for innovative, microbiome-targeted therapeutics. We finally propose a roadmap for future studies in the field for transforming critical care from its current isolated focus on the host to a more personalized paradigm addressing both human and microbial contributions to critical illness.
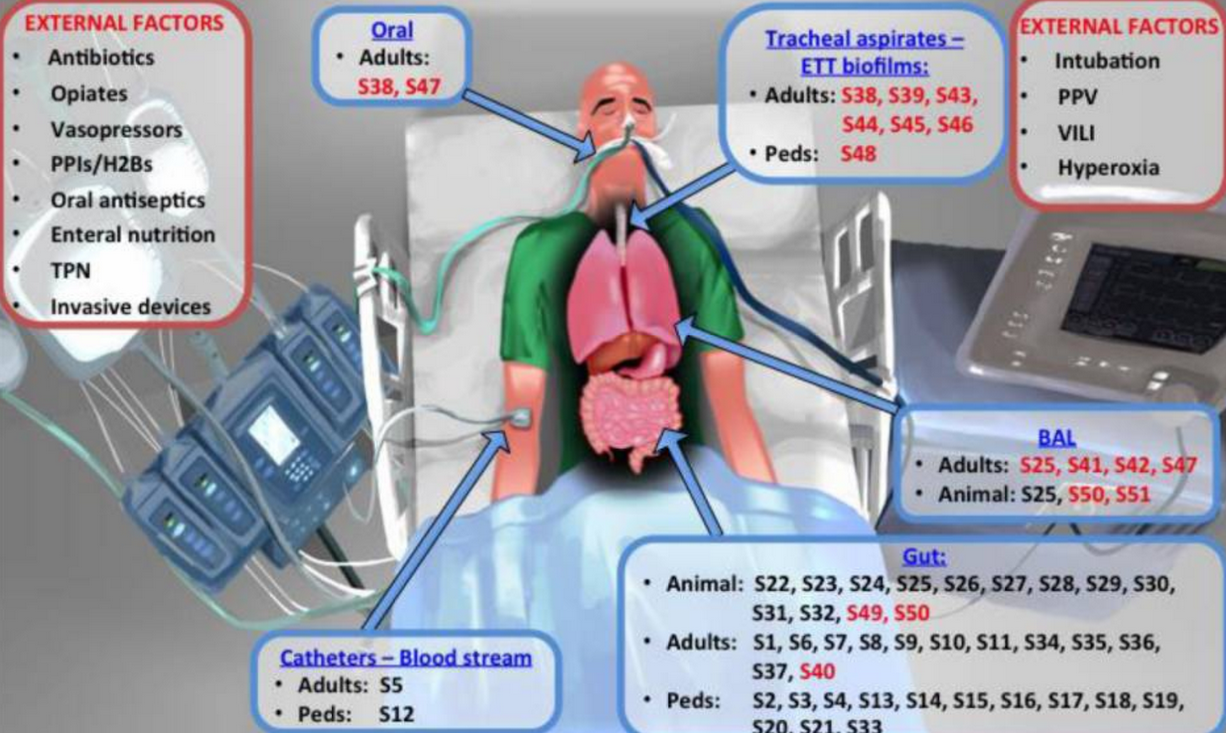
Factors that may alter the microbiome in the ICU (shown in red boxes) and references of reviewed microbiome studies according to body site sampled and study subjects (animal, adult or pediatric patients) (shown in blue boxes). Reference numbers (prefaced by “S”) correspond to the reference list provided in the Online Data Supplement. References for studies in sepsis are presented in black font and for studies in acute respiratory failure in red font.
Abbreviations: ETT: endotracheal tube; PPIs: proton-pump inhibitors; H2B: histamine-2 receptor blockers; TPN: total parenteral nutrition; PPV: positive pressure ventilation; VILI: ventilator-induced lung injury; Peds: pediatrics
The advent of molecular, culture-independent techniques to study micro-organisms revealed that the human host harbors approximately 40 trillion microbes, including bacteria and their phages, viruses, fungi and archaea. These microbes, organized in complex communities and contributing an enormous amount of genomic information, are clearly important, yet their roles are largely uncharacterized. An exponentially growing body of literature explores the role of the microbiome across a vast array of human pathologies, while the microbiome in critical care has not been studied as extensively. We have just begun to explore how microbiota perturbations (dysbiosis) are involved in the development, evolution and outcome of critical illness, and such microbiome research in patients in intensive care units (ICUs) holds tremendous potential. In the short-term, molecular techniques may allow us to provide more timely, accurate and personalized management of infections compared to our current practice directed by traditional microbial cultures. With deeper understanding of host-microbe interactions, microbiome research may reveal new targets for groundbreaking therapeutics for inflammatory syndromes, such as sepsis and the acute respiratory distress syndrome (ARDS), since efforts so far to modify host responses (without considering their microbial counterparts) have not delivered any efficacious therapies. In this article, we review the current state of knowledge on dysbiosis with critical illness and we also discuss important research challenges and strategies to move the field forward. We also provide a synopsis of available microbiome evidence for two common and serious clinical syndromes requiring care in general ICUs- sepsis and acute respiratory failure...Read full article here.
https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5328797/
